Drug-resistantbacteria was discovered in the technician’s garden
by admin
Lassocidin: A peptide-based antibiotic that kills pathogens using a lasso-shaped knot
Researchers have discovered a new antibiotic molecule that targets a broad range of disease-causing bacteria — even strains resistant to commercial drugs — and is not toxic to human cells1.
The molecule was discovered in the garden of a laboratory technician. The discovery shows that “there is terrifically interesting stuff hiding in plain sight”, says Kim Lewis, a microbiologist at Northeastern University in Boston, Massachusetts, who was not involved in the research. It’s good to know that they know what to look for.
The search for new antibiotics is necessary because bacteria acquire resistance to existing drugs with continued use. By 2020, the number of deaths associated with antimicrobial drugs could increase to 1.9 million.
Wright and his colleagues set out to find microbes that have developed previously unknown tricks to kill pathogens. They collected soil samples in Petri dishes with growth medium and stored them for a year. The researchers then exposed the microbes from these samples to Escherichia coli, a common gut bacterium that can cause serious disease.
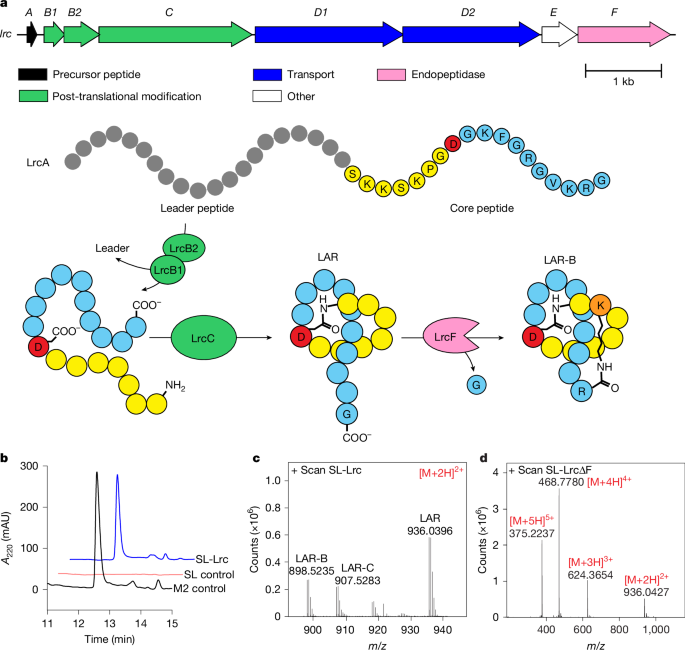
A molecule produced by the bacterium is a group of peptides, which forms a lasso-shaped knot. These peptides are known for being robust and can probably even survive being digested. Wright says the structure is compact and robust.
The molecule, which the researchers named lariocidin, binds to the ribosome and also to transfer RNA, which supplies the ribosome with the amino-acid building blocks it needs to string together peptide chains. By doing so, it prevents the genetic code from being read correctly, and also corrupts the code, jumbling the output. Ultimately, this means that the ribosome produces incorrect peptides, some of which probably end up being toxic to the bacterium and killing it, says Lewis. The pathogens haven’t developed resistance to the antibiotic because it uses a different mode of action.
E, Gavrish and some other people. Lassomycin has been shown to kill Mycobacterium tuberculosis by targeting ClpC1P1P2. The word Chem. means “chemicals.” Biol. 21, 509–518 (2014).
Mukhopadhyay, J., Sineva, E., Knight, J., Levy, R. M. & Ebright, R. H. Antibacterial peptide microcin J25 inhibits transcription by binding within and obstructing the RNA polymerase secondary channel. It’s known as the “mol.” Apparently it’s a scientific word. The Cell. 14, 739–751 was published in 2004.
Different modes of action of ribosome-targeting antibiotics and mechanisms of resistance are discussed. Annu. Rev. Biochem. 87, 451–478 (2018).
Sieber, S. A. & Marahiel, M. A. Molecular mechanisms underlying nonribosomal peptide synthesis: Approaches to new antibiotics. The Chem. Rev. 105 was published in 2005.
P. G., Arnison, and other people have worked on the project. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. They did an item called “pub.” Rep. 30, 108–160 (2013).
Resistance to seven classes ofprotein synthesis inhibitors can be achieved by the expression of the mlr operon. Antimicrob. Agents Chemother. 52, 1703–1712 (2008).
An experimental study of the role of rRNA modification in genetic assembly and drug resistance of Acinetobacter baumannii. Ann. Microbiol. 49, kuad042 (2019)
Cook, M. A. Lessons from assembling a microbial natural product and pre-fractionated extract library in an academic laboratory. J. Ind. Microbiol. There is a sugar molecule called Biotechnol. 50, kuad042.
Quan, S., Skovgaard, O., McLaughlin, R. E., Buurman, E. T. & Squires, C. L. Markerless Escherichia coli rrn deletion strains for genetic determination of ribosomal binding sites. G3 5, 2555–2557 (2015).
Structural insights into the role of rRNA modifications in ribosome assembly. Nat. Struct. The monograph of the Biol. 22, 342–344, was published in 2015.
The paper was written by Y.S. et al. Negamycin interferes with decoding and translocation by simultaneous interaction with rRNA and tRNA. Mol. Cell 56, 541–550 (2014).
Mistranslation of twocomponent system activation is associated with antibiotic-mediated cell death. Cell 135, 679–695.
Carson, D. V., Juarez, R. J., Do, T., Yang, Z. J. & Link, A. J. Antimicrobial lasso peptide cloacaenodin utilizes a unique TonB-dependent transporter to access susceptible bacteria. The substance is calledACS Chem. Biol. 19, 981–991 (2024).
Miller, S., Goy, K., She, R., Spellberg, B. & Luna, B. Antimicrobial susceptibility testing performed in RPMI 1640 reveals azithromycin efficacy against carbapenem-resistant Acinetobacter baumannii and predicts in vivo outcomes in Galleria mellonella. Antimicrob. The chemical agents are called Chemother.
Luna, B. et al. A nutrient-limited screen unmasks rifabutin hyperactivity for extensively drug-resistant Acinetobacter baumannii. Nat. Microbiol. There are five entries in this series, which will conclude in 2020.
Farha, M. A., French, S., Stokes, J. M. & Brown, E. D. Bicarbonate alters bacterial susceptibility to antibiotics by targeting the proton motive force. ACS Infect. Dis. 4, 382–390 (2018).
Source: A broad-spectrum lasso peptide antibiotic targeting the bacterial ribosome
Automatic assembly of the genomes of bacterial ribosomes: a comparative study of hybrid approaches by Chen, Z. and Yang, K. et al
Some hybrid assembly approaches for analysis of the genomes ofbacteria were compared by Chen, Z. It was the 21stBMC Genomics in 2020.
Blin, K. et al. AntiSMASH is a revision of predictions for detection, regulation, chemical structures and visualization. Nucleic Acids Res. 51, W46–w50 (2023).
C. L. M., and Y.-H. clinker are authors of automatic generation of gene cluster comparison figures. Bioinformatics 37, 2473–2475 (2021).
Jalview version 2-a multiple sequence alignment editor and workbench was written by A. M. and J. B. Bioinformatics 25, 1189–1191 (2009).
Bai, C. and others. Using a precise design of universal synthetic modular regulatory elements to unlock the natural products in Streptomyces. Proc. Natl Acad. The USA was covered inSci.USA 112, 12181–12186
Hong, H.J., Hutchings, M. I., Hill, L. M., and Buttner, M. J. The role of the novel Fem protein VanK in vancomycin resistance in Streptomyces coelicolor. J. Biol. Chem. 280 was published in 2005.
Deatherage and J. E. identified mutations in laboratory-evolved microbes from next-generation sequencing data. The methods are Mol. Biol. 1151, 165 and 186 were added in June.
Source: A broad-spectrum lasso peptide antibiotic targeting the bacterial ribosome
Towards a Structural Understanding of the Ribosome Function from the Structure of Non-Arrested ribosome-Nascent Chain Complexes
Syroegin, E. A., Aleksandrova, E. V. & Polikanov, Y. S. Insights into the ribosome function from the structures of non-arrested ribosome–nascent chain complexes. Nat. Chem. 15, 143–153 (2023).
Chen, C. W. et al. Structural insights into the mechanism of overcoming Erm-mediated resistance. Commun. 14, 4196.
Researchers have discovered a peptide-based antibiotic molecule that targets a broad range of disease-causing bacteria and is not toxic to human cells. The molecule, namedlariocidin, binds to the ribosome and also transfer RNA, supplies ribosome with aminoacid building blocks it needs to string together peptide chains. Ultimately, this means that ribosome produces incorrect peptides, some of probably end up being toxic to the bacterium.